Clustering and integration workflow
Songqi Duan
duan@songqi.org Source:vignettes/clustering.Rmd
clustering.RmdThis article covers the core unsupervised-analysis stage in Shennong using the built-in PBMC examples. The workflow shows how to move from a preprocessed Seurat object to:
- single-sample clustering
- multi-sample Harmony integration
- optional rare-aware feature selection before PCA/integration
- pre/post integration embedding comparison
- cluster visualization on the integrated object
Downstream metrics, composition, and interpretation now have their own dedicated articles, but a few summary outputs remain here so the integration workflow is visible end to end.
To keep R CMD check deterministic, the analysis is only
evaluated when SHENNONG_RUN_VIGNETTES=true or during
pkgdown builds.
Shennong also ships bundled human and mouse GENCODE gene annotations,
so sn_filter_genes() can combine expression-threshold
filtering with annotation-based retention such as
gene_class = "coding" or exact gene_type
subsets.
Inspect bundled signatures
Shennong now ships a tree-structured signature catalog derived from
SignatuR and stored as package data. You can inspect the
available signature paths and retrieve a specific signature either by
short alias or by full tree path.
signature_catalog <- sn_list_signatures(species = "human")
head(signature_catalog[, c("path", "n_genes")], 6)
#> # A tibble: 6 × 2
#> path n_genes
#> <chr> <int>
#> 1 Blocklists/Pseudogenes 12600
#> 2 Blocklists/Non-coding 7783
#> 3 Programs/HeatShock 97
#> 4 Programs/cellCycle.G1S 42
#> 5 Programs/cellCycle.G2M 52
#> 6 Programs/IFN 107
mito_genes <- sn_get_signatures(
species = "human",
category = c("mito", "Compartments/Ribo")
)
length(mito_genes)
#> [1] 205Data loaded for the workflow
knitr::kable(cell_summary, digits = 0)| dataset | cells | genes | clusters |
|---|---|---|---|
| pbmc1k | 1141 | 24986 | 11 |
| pbmc3k | 2671 | 19971 | NA |
| merged_unintegrated | 3812 | 25597 | 14 |
| merged_integrated | 3812 | 25597 | 11 |
The merged analysis is intentionally run twice: once without batch correction and once with Harmony integration. This makes the before/after comparison explicit instead of only showing the corrected embedding.
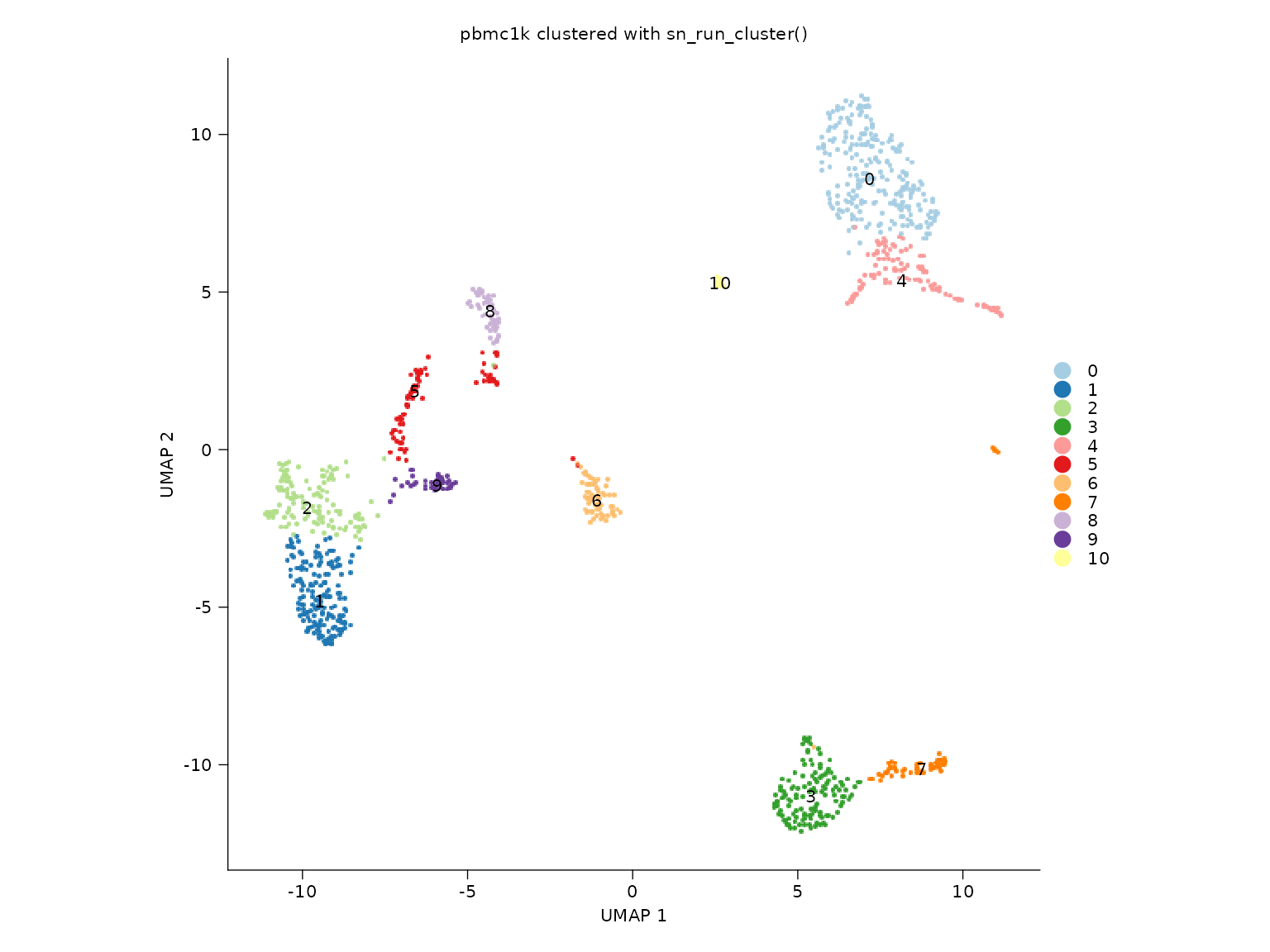
Single-sample clustering
sn_plot_dim(
object = pbmc1k_clustered,
reduction = "umap",
group_by = "seurat_clusters",
label = TRUE,
show_legend = FALSE,
title = "pbmc1k clustered with sn_run_cluster()"
)
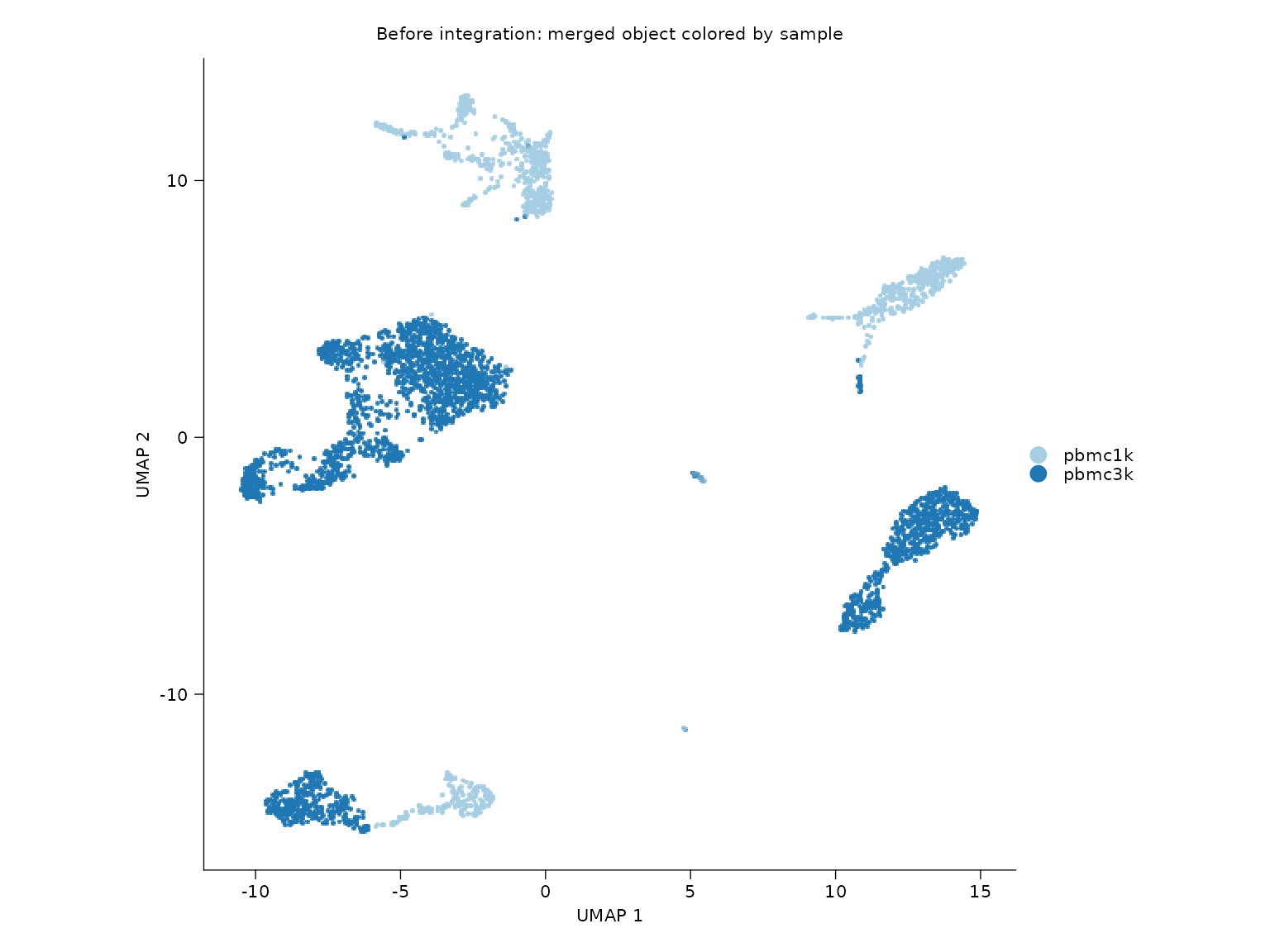
Before and after Harmony integration
sn_plot_dim(
object = pbmc_unintegrated,
reduction = "umap",
group_by = "sample",
title = "Before integration: merged object colored by sample"
)
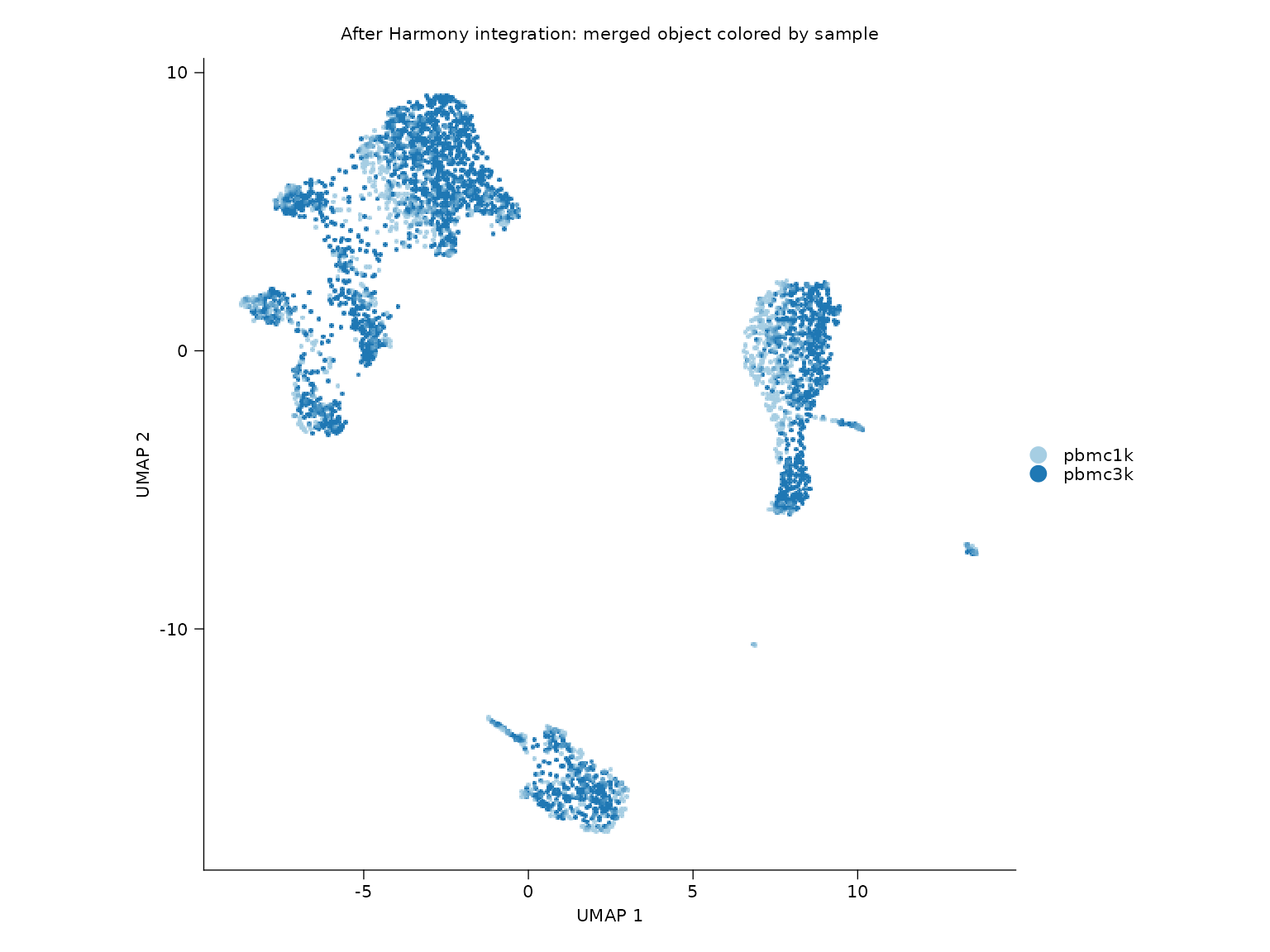
sn_plot_dim(
object = pbmc_integrated,
reduction = "umap",
group_by = "sample",
title = "After Harmony integration: merged object colored by sample"
)
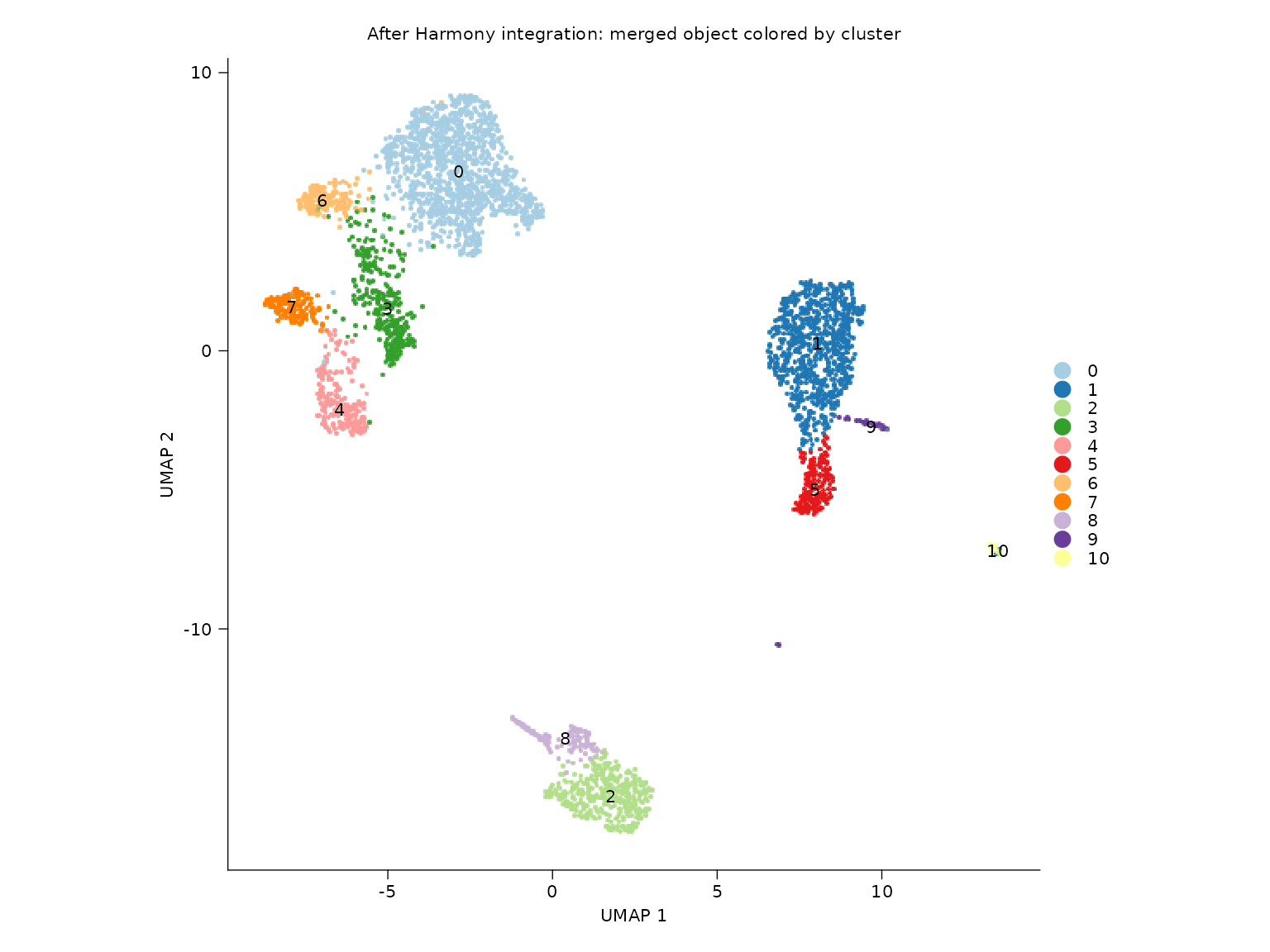
sn_plot_dim(
object = pbmc_integrated,
reduction = "umap",
group_by = "seurat_clusters",
label = TRUE,
show_legend = FALSE,
title = "After Harmony integration: merged object colored by cluster"
)
Cluster composition by sample
knitr::kable(dplyr::slice_head(composition_tbl, n = 12), digits = 2)| sample | seurat_clusters | proportion |
|---|---|---|
| pbmc1k | 0 | 27.61 |
| pbmc1k | 1 | 27.96 |
| pbmc1k | 10 | 1.31 |
| pbmc1k | 2 | 11.31 |
| pbmc1k | 3 | 5.96 |
| pbmc1k | 4 | 6.22 |
| pbmc1k | 5 | 2.28 |
| pbmc1k | 6 | 3.86 |
| pbmc1k | 7 | 6.13 |
| pbmc1k | 8 | 5.17 |
| pbmc1k | 9 | 2.19 |
| pbmc3k | 0 | 36.62 |
sn_plot_barplot(
composition_tbl,
x = sample,
y = proportion,
fill = seurat_clusters
) +
labs(
title = "Cluster composition across pbmc1k and pbmc3k",
x = NULL,
y = "Cells (%)",
fill = "Cluster"
) +
theme_minimal(base_size = 12)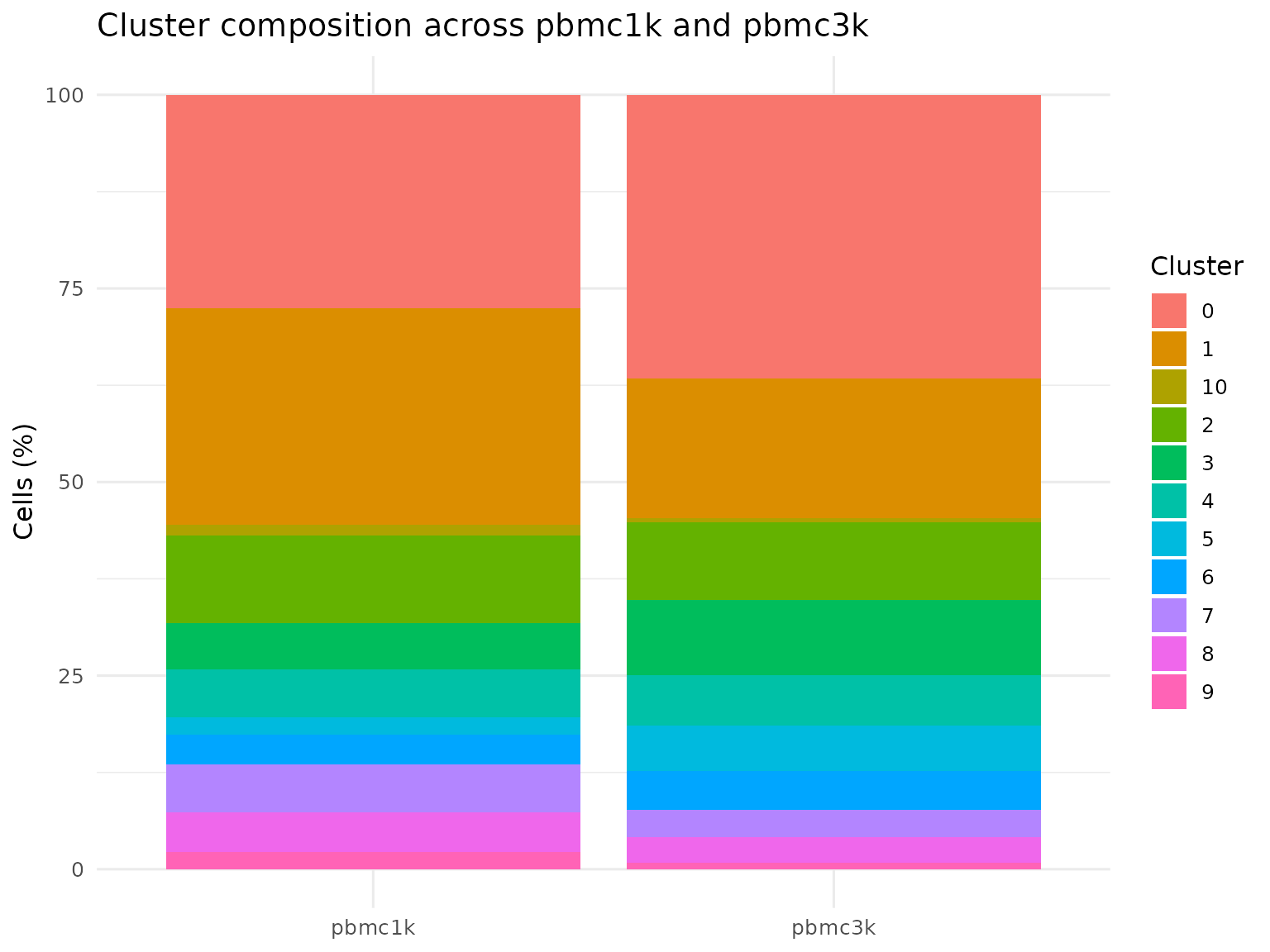
Integration quality summary with LISI
knitr::kable(lisi_summary, digits = 3)| state | min | q1 | median | mean | q3 | max |
|---|---|---|---|---|---|---|
| after_integration | 1.002 | 1.212 | 1.491 | 1.517 | 1.837 | 2 |
| before_integration | 1.000 | 1.000 | 1.000 | 1.024 | 1.000 | 2 |
Higher LISI values indicate better local sample mixing. In this two-sample example the theoretical maximum is 2, so the post-Harmony distribution should shift upward relative to the unintegrated baseline.
Multi-metric integration assessment
knitr::kable(assessment_summary, digits = 3)| metric | category | score | scaled_score | n_cells | source | note |
|---|---|---|---|---|---|---|
| batch_silhouette | batch_removal | 0.096 | 0.904 | 3812 | harmony | Global batch silhouette |
| batch_lisi | batch_removal | 1.517 | 0.517 | 3812 | harmony | |
| cluster_connectivity | structure | 1.000 | 1.000 | 3812 | RNA_nn | |
| pcr_batch | batch_removal | 0.007 | 0.929 | 3812 | harmony vs pca |
sn_assess_integration() combines local sample mixing,
PCR batch reduction, graph connectivity, isolated-label preservation,
cluster batch entropy, cluster purity, and difficult-group diagnostics
into one summary object while reusing the stored neighbor graph when
possible.
knitr::kable(challenging_groups, digits = 3)| seurat_clusters | n_cells | fraction_cells | median_neighbor_purity | mean_neighbor_purity | graph_connectivity | mean_silhouette | separation_score | challenge_score | rare_group | challenging_group |
|---|---|---|---|---|---|---|---|---|---|---|
| 9 | 48 | 0.013 | 0.956 | 0.836 | 1 | -0.041 | 0.812 | 0.188 | TRUE | FALSE |
| 8 | 146 | 0.038 | 0.969 | 0.903 | 1 | 0.102 | 0.840 | 0.160 | FALSE | FALSE |
| 3 | 329 | 0.086 | 1.000 | 0.886 | 1 | 0.058 | 0.843 | 0.157 | FALSE | FALSE |
| 0 | 1293 | 0.339 | 1.000 | 0.977 | 1 | 0.196 | 0.866 | 0.134 | FALSE | FALSE |
| 4 | 244 | 0.064 | 1.000 | 0.952 | 1 | 0.223 | 0.870 | 0.130 | FALSE | FALSE |
| 2 | 395 | 0.104 | 1.000 | 0.967 | 1 | 0.281 | 0.880 | 0.120 | FALSE | FALSE |
| 6 | 177 | 0.046 | 1.000 | 0.907 | 1 | 0.305 | 0.884 | 0.116 | FALSE | FALSE |
| 7 | 165 | 0.043 | 1.000 | 0.939 | 1 | 0.313 | 0.886 | 0.114 | FALSE | FALSE |
| 5 | 184 | 0.048 | 1.000 | 0.962 | 1 | 0.376 | 0.896 | 0.104 | FALSE | FALSE |
| 1 | 801 | 0.210 | 1.000 | 0.987 | 1 | 0.378 | 0.896 | 0.104 | FALSE | FALSE |
| 10 | 30 | 0.008 | 1.000 | 0.995 | 1 | 0.738 | 0.956 | 0.044 | TRUE | FALSE |
The challenging_groups table is especially useful for
surfacing rare groups or poorly separated populations that may not stand
out as isolated UMAP islands.
knitr::kable(integration_assessment$per_group$isolated_label_score, digits = 3)The isolated-label summary makes it easier to spot small populations that stay biologically distinct even when they do not appear as a visually isolated UMAP island after integration.
Rare-aware feature selection
When small populations are a concern, sn_run_cluster()
can append rare-aware features to the main HVG set before PCA and
Harmony. In this article the integrated object was run with
rare_feature_method = "gini", and the resulting rare-cell
scores can be inspected directly:
knitr::kable(head(rare_detection[order(rare_detection$rare_score, decreasing = TRUE), ], 10), digits = 3)| cell_id | method | rare_score | rare_cell | |
|---|---|---|---|---|
| 640 | pbmc1k_GCCGTGAGTCAGTCCG-1 | gini | 79.314 | TRUE |
| 838 | pbmc1k_TACCGGGTCCTCGATC-1 | gini | 78.042 | TRUE |
| 1040 | pbmc1k_TGTAACGGTTGCGTAT-1 | gini | 71.848 | TRUE |
| 148 | pbmc1k_AGCCAGCGTAGTTAGA-1 | gini | 69.761 | TRUE |
| 674 | pbmc1k_GGATCTAGTGCCTGCA-1 | gini | 69.457 | TRUE |
| 1093 | pbmc1k_TTCCTAAGTGACGTCC-1 | gini | 63.962 | TRUE |
| 801 | pbmc1k_GTGTTCCGTAGTTCCA-1 | gini | 61.457 | TRUE |
| 32 | pbmc1k_AACGAAAAGGTTGGTG-1 | gini | 52.102 | TRUE |
| 1089 | pbmc1k_TTCCACGAGGAAGTGA-1 | gini | 47.752 | TRUE |
| 192 | pbmc1k_AGTCACATCTCCCATG-1 | gini | 47.373 | TRUE |
Marker genes from the integrated object
knitr::kable(marker_tbl, digits = 3)| cluster | gene | avg_log2FC | p_val_adj |
|---|---|---|---|
| 0 | TSHZ2 | 5.214 | 0 |
| 0 | ENSG00000249806 | 4.826 | 0 |
| 0 | CMTM8 | 3.704 | 0 |
| 1 | S100A12 | 8.653 | 0 |
| 1 | FOLR3 | 8.427 | 0 |
| 1 | CLEC4E | 7.854 | 0 |
| 2 | TCL1A | 8.067 | 0 |
| 2 | IGHD | 6.823 | 0 |
| 2 | PCDH9 | 6.358 | 0 |
| 3 | GZMH | 4.361 | 0 |
| 3 | TRGC2 | 4.120 | 0 |
| 3 | CD8A | 3.721 | 0 |
| 4 | ENSG00000289191 | 7.495 | 0 |
| 4 | SH2D1B | 7.266 | 0 |
| 4 | GNLY | 7.205 | 0 |
| 5 | ENSG00000301038 | 8.430 | 0 |
| 5 | CKB | 7.015 | 0 |
| 5 | CDKN1C | 6.909 | 0 |
| 6 | LINC02446 | 5.616 | 0 |
| 6 | ENSG00000310107 | 4.854 | 0 |
| 6 | NRCAM | 4.647 | 0 |
| 7 | SLC4A10 | 7.660 | 0 |
| 7 | ENSG00000228033 | 7.646 | 0 |
| 7 | ADAM12 | 6.413 | 0 |
| 8 | SSPN | 8.885 | 0 |
| 8 | IGHA1 | 8.245 | 0 |
| 8 | IGHG3 | 7.907 | 0 |
| 9 | LINC01478 | 10.605 | 0 |
| 9 | FCER1A | 8.531 | 0 |
| 9 | DNASE1L3 | 8.426 | 0 |
| 10 | PF4V1 | 15.175 | 0 |
| 10 | CLDN5 | 13.563 | 0 |
| 10 | CMTM5 | 13.046 | 0 |
sn_plot_dot(
x = pbmc_integrated,
features = "top_markers",
de_name = "cluster_markers",
n = 3
)
Functional enrichment of marker genes
The table below shows the first non-empty GO Biological Process GSEA result found among the cluster marker sets. In this run, enrichment was computed from cluster 0.
knitr::kable(enrichment_tbl, digits = 4)| ID | Description | NES | p.adjust | |
|---|---|---|---|---|
| GO:0002250 | GO:0002250 | adaptive immune response | 2.2222 | 0 |
| GO:0042110 | GO:0042110 | T cell activation | 2.1480 | 0 |
| GO:0046649 | GO:0046649 | lymphocyte activation | 1.9937 | 0 |
| GO:0001775 | GO:0001775 | cell activation | 1.9338 | 0 |
| GO:0045321 | GO:0045321 | leukocyte activation | 1.9244 | 0 |
| GO:0006955 | GO:0006955 | immune response | 1.8855 | 0 |
| GO:1903131 | GO:1903131 | mononuclear cell differentiation | 2.0001 | 0 |
| GO:0002684 | GO:0002684 | positive regulation of immune system process | 1.9479 | 0 |
| GO:0002682 | GO:0002682 | regulation of immune system process | 1.8416 | 0 |
| GO:0002768 | GO:0002768 | immune response-regulating cell surface receptor signaling pathway | 2.1336 | 0 |
sessioninfo::session_info()
#> ─ Session info ───────────────────────────────────────────────────────────────
#> setting value
#> version R version 4.5.3 (2026-03-11)
#> os Ubuntu 24.04.3 LTS
#> system x86_64, linux-gnu
#> ui X11
#> language en
#> collate C.UTF-8
#> ctype C.UTF-8
#> tz UTC
#> date 2026-03-26
#> pandoc 3.1.11 @ /opt/hostedtoolcache/pandoc/3.1.11/x64/ (via rmarkdown)
#> quarto NA
#>
#> ─ Packages ───────────────────────────────────────────────────────────────────
#> package * version date (UTC) lib source
#> abind 1.4-8 2024-09-12 [1] CRAN (R 4.5.3)
#> AnnotationDbi 1.72.0 2025-10-29 [1] Bioconduc~
#> ape 5.8-1 2024-12-16 [1] CRAN (R 4.5.3)
#> aplot 0.2.9 2025-09-12 [1] CRAN (R 4.5.3)
#> Biobase 2.70.0 2025-10-29 [1] Bioconduc~
#> BiocGenerics 0.56.0 2025-10-29 [1] Bioconduc~
#> BiocParallel 1.44.0 2025-10-29 [1] Bioconduc~
#> Biostrings 2.78.0 2025-10-29 [1] Bioconduc~
#> bit 4.6.0 2025-03-06 [1] CRAN (R 4.5.3)
#> bit64 4.6.0-1 2025-01-16 [1] CRAN (R 4.5.3)
#> blob 1.3.0 2026-01-14 [1] CRAN (R 4.5.3)
#> bslib 0.10.0 2026-01-26 [1] CRAN (R 4.5.3)
#> cachem 1.1.0 2024-05-16 [1] CRAN (R 4.5.3)
#> catplot 0.1.0 2026-03-19 [1] Github (catplot/catplot@0fc2344)
#> cli 3.6.5 2025-04-23 [1] CRAN (R 4.5.3)
#> cluster 2.1.8.2 2026-02-05 [3] CRAN (R 4.5.3)
#> clusterProfiler 4.18.4 2025-12-15 [1] any (@4.18.4)
#> codetools 0.2-20 2024-03-31 [3] CRAN (R 4.5.3)
#> cowplot 1.2.0 2025-07-07 [1] CRAN (R 4.5.3)
#> crayon 1.5.3 2024-06-20 [1] CRAN (R 4.5.3)
#> curl 7.0.0 2025-08-19 [1] CRAN (R 4.5.3)
#> data.table 1.18.2.1 2026-01-27 [1] CRAN (R 4.5.3)
#> DBI 1.3.0 2026-02-25 [1] CRAN (R 4.5.3)
#> deldir 2.0-4 2024-02-28 [1] CRAN (R 4.5.3)
#> desc 1.4.3 2023-12-10 [1] CRAN (R 4.5.3)
#> digest 0.6.39 2025-11-19 [1] CRAN (R 4.5.3)
#> DOSE 4.4.0 2025-10-29 [1] Bioconduc~
#> dotCall64 1.2 2024-10-04 [1] CRAN (R 4.5.3)
#> dplyr * 1.2.0 2026-02-03 [1] CRAN (R 4.5.3)
#> enrichplot 1.30.5 2026-03-02 [1] Bioconduc~
#> evaluate 1.0.5 2025-08-27 [1] CRAN (R 4.5.3)
#> farver 2.1.2 2024-05-13 [1] CRAN (R 4.5.3)
#> fastDummies 1.7.5 2025-01-20 [1] CRAN (R 4.5.3)
#> fastmap 1.2.0 2024-05-15 [1] CRAN (R 4.5.3)
#> fastmatch 1.1-8 2026-01-17 [1] CRAN (R 4.5.3)
#> fgsea 1.36.2 2026-01-05 [1] Bioconduc~
#> fitdistrplus 1.2-6 2026-01-24 [1] CRAN (R 4.5.3)
#> fontBitstreamVera 0.1.1 2017-02-01 [1] CRAN (R 4.5.3)
#> fontLiberation 0.1.0 2016-10-15 [1] CRAN (R 4.5.3)
#> fontquiver 0.2.1 2017-02-01 [1] CRAN (R 4.5.3)
#> fs 2.0.1 2026-03-24 [1] CRAN (R 4.5.3)
#> future * 1.70.0 2026-03-14 [1] CRAN (R 4.5.3)
#> future.apply 1.20.2 2026-02-20 [1] CRAN (R 4.5.3)
#> gdtools 0.5.0 2026-02-09 [1] CRAN (R 4.5.3)
#> generics 0.1.4 2025-05-09 [1] CRAN (R 4.5.3)
#> ggforce 0.5.0 2025-06-18 [1] CRAN (R 4.5.3)
#> ggfun 0.2.0 2025-07-15 [1] CRAN (R 4.5.3)
#> ggiraph 0.9.6 2026-02-21 [1] CRAN (R 4.5.3)
#> ggnewscale 0.5.2 2025-06-20 [1] CRAN (R 4.5.3)
#> ggplot2 * 4.0.2 2026-02-03 [1] CRAN (R 4.5.3)
#> ggplotify 0.1.3 2025-09-20 [1] CRAN (R 4.5.3)
#> ggrepel 0.9.8 2026-03-17 [1] CRAN (R 4.5.3)
#> ggridges 0.5.7 2025-08-27 [1] CRAN (R 4.5.3)
#> ggtangle 0.1.1 2026-01-16 [1] CRAN (R 4.5.3)
#> ggtree 4.0.5 2026-03-17 [1] Bioconduc~
#> globals 0.19.1 2026-03-13 [1] CRAN (R 4.5.3)
#> glue 1.8.0 2024-09-30 [1] CRAN (R 4.5.3)
#> GO.db 3.22.0 2026-03-19 [1] Bioconductor
#> goftest 1.2-3 2021-10-07 [1] CRAN (R 4.5.3)
#> GOSemSim 2.36.0 2025-10-29 [1] Bioconduc~
#> gridExtra 2.3 2017-09-09 [1] CRAN (R 4.5.3)
#> gridGraphics 0.5-1 2020-12-13 [1] CRAN (R 4.5.3)
#> gson 0.1.0 2023-03-07 [1] CRAN (R 4.5.3)
#> gtable 0.3.6 2024-10-25 [1] CRAN (R 4.5.3)
#> harmony 2.0.0 2026-03-24 [1] Github (immunogenomics/harmony@3617c00)
#> hdf5r 1.3.12 2025-01-20 [1] any (@1.3.12)
#> HGNChelper 0.8.15 2024-11-16 [1] any (@0.8.15)
#> htmltools 0.5.9 2025-12-04 [1] CRAN (R 4.5.3)
#> htmlwidgets 1.6.4 2023-12-06 [1] CRAN (R 4.5.3)
#> httpuv 1.6.17 2026-03-18 [1] CRAN (R 4.5.3)
#> httr 1.4.8 2026-02-13 [1] CRAN (R 4.5.3)
#> ica 1.0-3 2022-07-08 [1] CRAN (R 4.5.3)
#> igraph 2.2.2 2026-02-12 [1] CRAN (R 4.5.3)
#> IRanges 2.44.0 2025-10-29 [1] Bioconduc~
#> irlba 2.3.7 2026-01-30 [1] CRAN (R 4.5.3)
#> jquerylib 0.1.4 2021-04-26 [1] CRAN (R 4.5.3)
#> jsonlite 2.0.0 2025-03-27 [1] CRAN (R 4.5.3)
#> KEGGREST 1.50.0 2025-10-29 [1] Bioconduc~
#> KernSmooth 2.23-26 2025-01-01 [3] CRAN (R 4.5.3)
#> knitr * 1.51 2025-12-20 [1] CRAN (R 4.5.3)
#> labeling 0.4.3 2023-08-29 [1] CRAN (R 4.5.3)
#> later 1.4.8 2026-03-05 [1] CRAN (R 4.5.3)
#> lattice 0.22-9 2026-02-09 [3] CRAN (R 4.5.3)
#> lazyeval 0.2.2 2019-03-15 [1] CRAN (R 4.5.3)
#> lifecycle 1.0.5 2026-01-08 [1] CRAN (R 4.5.3)
#> limma 3.66.0 2025-10-29 [1] Bioconduc~
#> lisi 1.0 2026-03-19 [1] Github (immunogenomics/lisi@a917556)
#> listenv 0.10.1 2026-03-10 [1] CRAN (R 4.5.3)
#> lmtest 0.9-40 2022-03-21 [1] CRAN (R 4.5.3)
#> logger 0.4.1 2025-09-11 [1] CRAN (R 4.5.3)
#> magrittr 2.0.4 2025-09-12 [1] CRAN (R 4.5.3)
#> MASS 7.3-65 2025-02-28 [3] CRAN (R 4.5.3)
#> Matrix 1.7-4 2025-08-28 [3] CRAN (R 4.5.3)
#> matrixStats 1.5.0 2025-01-07 [1] CRAN (R 4.5.3)
#> memoise 2.0.1 2021-11-26 [1] CRAN (R 4.5.3)
#> mime 0.13 2025-03-17 [1] CRAN (R 4.5.3)
#> miniUI 0.1.2 2025-04-17 [1] CRAN (R 4.5.3)
#> nlme 3.1-168 2025-03-31 [3] CRAN (R 4.5.3)
#> org.Hs.eg.db 3.22.0 2026-03-19 [1] bioc (@3.22.0)
#> otel 0.2.0 2025-08-29 [1] CRAN (R 4.5.3)
#> parallelly 1.46.1 2026-01-08 [1] CRAN (R 4.5.3)
#> patchwork 1.3.2 2025-08-25 [1] CRAN (R 4.5.3)
#> pbapply 1.7-4 2025-07-20 [1] CRAN (R 4.5.3)
#> pillar 1.11.1 2025-09-17 [1] CRAN (R 4.5.3)
#> pkgconfig 2.0.3 2019-09-22 [1] CRAN (R 4.5.3)
#> pkgdown 2.2.0 2025-11-06 [1] any (@2.2.0)
#> plotly 4.12.0 2026-01-24 [1] CRAN (R 4.5.3)
#> plyr 1.8.9 2023-10-02 [1] CRAN (R 4.5.3)
#> png 0.1-9 2026-03-15 [1] CRAN (R 4.5.3)
#> polyclip 1.10-7 2024-07-23 [1] CRAN (R 4.5.3)
#> progressr 0.18.0 2025-11-06 [1] CRAN (R 4.5.3)
#> promises 1.5.0 2025-11-01 [1] CRAN (R 4.5.3)
#> purrr 1.2.1 2026-01-09 [1] CRAN (R 4.5.3)
#> qvalue 2.42.0 2025-10-29 [1] Bioconduc~
#> R.methodsS3 1.8.2 2022-06-13 [1] CRAN (R 4.5.3)
#> R.oo 1.27.1 2025-05-02 [1] CRAN (R 4.5.3)
#> R.utils 2.13.0 2025-02-24 [1] CRAN (R 4.5.3)
#> R6 2.6.1 2025-02-15 [1] CRAN (R 4.5.3)
#> ragg 1.5.2 2026-03-23 [1] CRAN (R 4.5.3)
#> RANN 2.6.2 2024-08-25 [1] CRAN (R 4.5.3)
#> rappdirs 0.3.4 2026-01-17 [1] CRAN (R 4.5.3)
#> RColorBrewer 1.1-3 2022-04-03 [1] CRAN (R 4.5.3)
#> Rcpp 1.1.1 2026-01-10 [1] CRAN (R 4.5.3)
#> RcppAnnoy 0.0.23 2026-01-12 [1] CRAN (R 4.5.3)
#> RcppHNSW 0.6.0 2024-02-04 [1] CRAN (R 4.5.3)
#> reshape2 1.4.5 2025-11-12 [1] CRAN (R 4.5.3)
#> reticulate 1.45.0 2026-02-13 [1] CRAN (R 4.5.3)
#> RhpcBLASctl 0.23-42 2023-02-11 [1] CRAN (R 4.5.3)
#> rio 1.2.4 2025-09-26 [1] any (@1.2.4)
#> rlang 1.1.7 2026-01-09 [1] CRAN (R 4.5.3)
#> rmarkdown 2.31 2026-03-26 [1] CRAN (R 4.5.3)
#> ROCR 1.0-12 2026-01-23 [1] CRAN (R 4.5.3)
#> RSpectra 0.16-2 2024-07-18 [1] CRAN (R 4.5.3)
#> RSQLite 2.4.6 2026-02-06 [1] CRAN (R 4.5.3)
#> Rtsne 0.17 2023-12-07 [1] CRAN (R 4.5.3)
#> S4Vectors 0.48.0 2025-10-29 [1] Bioconduc~
#> S7 0.2.1 2025-11-14 [1] CRAN (R 4.5.3)
#> sass 0.4.10 2025-04-11 [1] CRAN (R 4.5.3)
#> scales 1.4.0 2025-04-24 [1] CRAN (R 4.5.3)
#> scattermore 1.2 2023-06-12 [1] CRAN (R 4.5.3)
#> scatterpie 0.2.6 2025-09-12 [1] CRAN (R 4.5.3)
#> sctransform 0.4.3 2026-01-10 [1] CRAN (R 4.5.3)
#> Seqinfo 1.0.0 2025-10-29 [1] Bioconduc~
#> sessioninfo 1.2.3 2025-02-05 [1] any (@1.2.3)
#> Seurat * 5.4.0 2025-12-14 [1] any (@5.4.0)
#> SeuratObject * 5.3.0 2025-12-12 [1] CRAN (R 4.5.3)
#> Shennong * 0.1.2 2026-03-26 [1] local
#> shiny 1.13.0 2026-02-20 [1] CRAN (R 4.5.3)
#> sp * 2.2-1 2026-02-13 [1] CRAN (R 4.5.3)
#> spam 2.11-3 2026-01-08 [1] CRAN (R 4.5.3)
#> spatstat.data 3.1-9 2025-10-18 [1] CRAN (R 4.5.3)
#> spatstat.explore 3.8-0 2026-03-22 [1] CRAN (R 4.5.3)
#> spatstat.geom 3.7-3 2026-03-23 [1] CRAN (R 4.5.3)
#> spatstat.random 3.4-5 2026-03-22 [1] CRAN (R 4.5.3)
#> spatstat.sparse 3.1-0 2024-06-21 [1] CRAN (R 4.5.3)
#> spatstat.univar 3.1-7 2026-03-18 [1] CRAN (R 4.5.3)
#> spatstat.utils 3.2-2 2026-03-10 [1] CRAN (R 4.5.3)
#> splitstackshape 1.4.8.1 2026-03-21 [1] CRAN (R 4.5.3)
#> statmod 1.5.1 2025-10-09 [1] CRAN (R 4.5.3)
#> stringi 1.8.7 2025-03-27 [1] CRAN (R 4.5.3)
#> stringr 1.6.0 2025-11-04 [1] CRAN (R 4.5.3)
#> survival 3.8-6 2026-01-16 [3] CRAN (R 4.5.3)
#> systemfonts 1.3.2 2026-03-05 [1] CRAN (R 4.5.3)
#> tensor 1.5.1 2025-06-17 [1] CRAN (R 4.5.3)
#> textshaping 1.0.5 2026-03-06 [1] CRAN (R 4.5.3)
#> tibble 3.3.1 2026-01-11 [1] CRAN (R 4.5.3)
#> tictoc 1.2.1 2024-03-18 [1] CRAN (R 4.5.3)
#> tidydr 0.0.6 2025-07-25 [1] CRAN (R 4.5.3)
#> tidyr 1.3.2 2025-12-19 [1] CRAN (R 4.5.3)
#> tidyselect 1.2.1 2024-03-11 [1] CRAN (R 4.5.3)
#> tidytree 0.4.7 2026-01-08 [1] CRAN (R 4.5.3)
#> treeio 1.34.0 2025-10-30 [1] Bioconduc~
#> tweenr 2.0.3 2024-02-26 [1] CRAN (R 4.5.3)
#> utf8 1.2.6 2025-06-08 [1] CRAN (R 4.5.3)
#> uwot 0.2.4 2025-11-10 [1] CRAN (R 4.5.3)
#> vctrs 0.7.2 2026-03-21 [1] CRAN (R 4.5.3)
#> viridisLite 0.4.3 2026-02-04 [1] CRAN (R 4.5.3)
#> withr 3.0.2 2024-10-28 [1] CRAN (R 4.5.3)
#> xfun 0.57 2026-03-20 [1] CRAN (R 4.5.3)
#> xtable 1.8-8 2026-02-22 [1] CRAN (R 4.5.3)
#> XVector 0.50.0 2025-10-29 [1] Bioconduc~
#> yaml 2.3.12 2025-12-10 [1] CRAN (R 4.5.3)
#> yulab.utils 0.2.4 2026-02-02 [1] CRAN (R 4.5.3)
#> zoo 1.8-15 2025-12-15 [1] CRAN (R 4.5.3)
#>
#> [1] /home/runner/work/_temp/Library
#> [2] /opt/R/4.5.3/lib/R/site-library
#> [3] /opt/R/4.5.3/lib/R/library
#> * ── Packages attached to the search path.
#>
#> ──────────────────────────────────────────────────────────────────────────────